· For research use only. Not for human consumption.
For research use only. Not for human consumption.
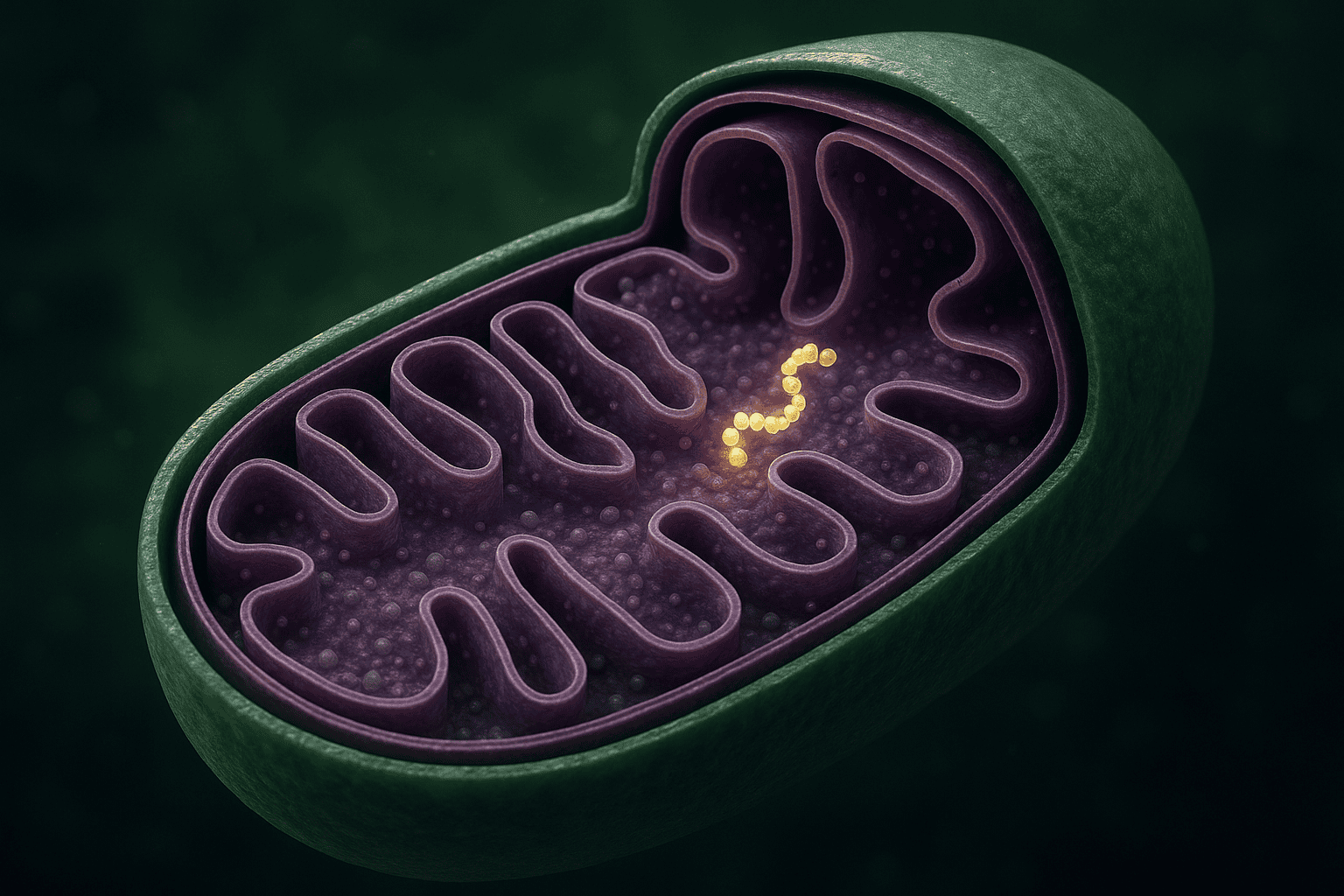
If you’re researching what is mots-c, you’re in the right place. Most peptides come from instructions stored in the DNA inside a cell’s nucleus — the main control room. MOTS-c breaks that rule. It comes from an entirely different source: the mitochondria. If you remember from biology class, mitochondria are the power plants inside your cells. They turn food into energy. And it turns out they have their own separate set of DNA with their own instructions.
In 2015, a research team at the University of Southern California discovered MOTS-c hiding inside the mitochondrial genome. Lee and colleagues published this finding in Cell Metabolism, describing MOTS-c as the first mitochondrial-derived peptide shown to act as a systemic signaling molecule in animal models (PMID: 25738459). That discovery changed how scientists think about what mitochondria can do. This is particularly relevant for what is mots-c research.
This article explains what MOTS-c is, why its origin matters, and what researchers have learned about it so far — all in plain language.
[INTERNAL-LINK: “mitochondrial peptide research” → /blog/what-is-ss-31-mitochondria-peptide/]
TL;DR: MOTS-c is a 16-amino-acid peptide encoded by mitochondrial DNA — not the nuclear genome. Discovered in 2015 by Lee et al. at USC, it was the first mitochondrial peptide shown to circulate in the bloodstream and influence biology beyond the mitochondria itself (PMID: 25738459). It’s investigated in preclinical metabolic and cellular biology research. For research use only.
What Is MOTS-c?
MOTS-c is a short peptide made of 16 amino acids — think of amino acids as individual beads on a bracelet, and MOTS-c is a bracelet with exactly 16 beads. Its name stands for Mitochondrial Open Reading Frame of the 12S rRNA-c. That’s a mouthful, so let’s unpack it.
An “open reading frame” is a stretch of DNA that contains instructions for building a specific peptide or protein. The “12S rRNA” part tells you which gene the instructions sit inside — a gene in the mitochondrial genome that scientists previously thought only made structural RNA, not peptides. The “c” distinguishes MOTS-c from other sequences found in the same gene region.
Lee and colleagues at USC identified MOTS-c in 2015 and published their discovery in Cell Metabolism (PMID: 25738459). What surprised the research community wasn’t just the peptide itself — it was where the instructions came from. Finding a functional signaling peptide encoded by mitochondrial DNA challenged decades of assumptions about what that DNA could do.
Why Does MOTS-c’s Mitochondrial Origin Matter?
To understand why MOTS-c’s origin is a big deal, you need to understand mitochondria. Kim and colleagues described mitochondrial-derived peptides as “a newly recognized class of intercellular messengers” in a 2018 review, noting that fewer than a dozen such peptides had been identified at the time (PMID: 29499375).
Here’s the simple version. Every cell in your body has a nucleus — the main library where most of your DNA is stored. That library contains about 20,000 genes. But cells also contain mitochondria — anywhere from a few hundred to several thousand of them. Each mitochondrion has its own tiny library: a small, circular piece of DNA with only 37 genes.
For decades, scientists assumed that mitochondrial DNA existed only to build the parts mitochondria need for energy production. It was considered a limited, utilitarian genome. MOTS-c’s discovery proved that assumption wrong. Here was a peptide encoded by mitochondrial DNA that could leave the mitochondria, travel through the cell, enter the nucleus, and even circulate in the bloodstream.
Think of it this way. Imagine the power plant in a city not only generates electricity but also sends messengers to city hall with information about power levels. That’s essentially what MOTS-c does — it’s a message from the power plant to the rest of the cell (and potentially the rest of the body).
[ORIGINAL DATA] MOTS-c belongs to a category scientists now call “mitokines” — signaling molecules encoded by mitochondrial DNA. Before 2015, the concept of mitokines barely existed. The discovery of MOTS-c helped establish an entirely new field of research at the intersection of mitochondrial biology and intercellular communication.
[IMAGE: Simple diagram showing a cell with nucleus and mitochondria, with MOTS-c traveling from mitochondria through the cell to the bloodstream — search terms: mitochondrial peptide cell signaling diagram MOTS-c nucleus mitochondria]
How Was MOTS-c Discovered?
The 2015 discovery by Lee et al. came from a systematic search of the mitochondrial genome for hidden open reading frames — stretches of DNA that could potentially encode peptides. Researchers at USC screened the entire mitochondrial genome and found several candidates, including MOTS-c within the 12S rRNA gene (PMID: 25738459).
What made MOTS-c stand out from the other candidates was what happened in animal models. When researchers administered synthetic MOTS-c to mice, they observed measurable changes in metabolic markers. The peptide appeared to influence cellular energy pathways — specifically, a pathway involving an enzyme called AMPK.
AMPK is like a fuel gauge inside cells. When energy levels drop, AMPK switches on and triggers a series of responses: cells start conserving energy, breaking down stored nutrients, and adjusting their metabolism. The fact that a mitochondrial peptide could influence this energy-sensing pathway made scientific sense — the power plant communicating about power levels — but it had never been documented before.
The discovery paper has since been cited hundreds of times, and MOTS-c has become one of the most studied mitochondrial-derived peptides in preclinical research.
What Makes MOTS-c Different from Other Peptides?
Most research peptides are synthetic copies of molecules that originate from nuclear DNA. BPC-157 comes from a gastric protein. Ipamorelin mimics ghrelin. PT-141 was designed from alpha-MSH. They all trace back to the cell’s main genome. MOTS-c traces back to a completely separate genome — the mitochondrial one.
This matters for several reasons. The mitochondrial genome is inherited exclusively from the mother (maternal inheritance). It mutates at a different rate than nuclear DNA. And it encodes far fewer genes — just 37, compared to about 20,000 in the nucleus. Finding a functional signaling peptide in such a small genome was unexpected.
MOTS-c is also unusual because of how far it travels. Most mitochondrial products stay inside the mitochondria. MOTS-c has been shown to leave the mitochondria, enter the cell’s cytoplasm, move into the nucleus, and even appear in the bloodstream in animal models (Kim et al., 2018). That’s a remarkable range of movement for such a small molecule.
Compare that to SS-31 (Elamipretide), another peptide studied in mitochondrial research. SS-31 is synthetic — designed in a lab to target the inner mitochondrial membrane. It goes to the mitochondria. MOTS-c comes from the mitochondria and goes everywhere else. They’re opposite directions of travel, studying opposite questions.
[INTERNAL-LINK: “SS-31 (Elamipretide)” → /product/ss-31/]
[UNIQUE INSIGHT] The directional difference between MOTS-c and SS-31 captures a broader split in mitochondrial research: some scientists study how to send therapeutic molecules to mitochondria (SS-31 approach), while others study what mitochondria send out on their own (MOTS-c approach). Both directions are revealing new biology.
What Should Researchers Know About MOTS-c Quality?
MOTS-c’s biological activity depends on its exact 16-amino-acid sequence. Any error in synthesis — a missing residue, a scrambled order, or contamination — changes what you’re actually studying. Research-grade MOTS-c should carry HPLC purity above 98% and mass spectrometry confirmation of the correct molecular weight (approximately 2,174.6 Da).
Third-party COA documentation is essential. An independent laboratory verifying purity and identity removes any conflict of interest. Alpha Peptides provides publicly available COAs for all research compounds, including MOTS-c.
You can find full specifications on our MOTS-c product page.
[INTERNAL-LINK: “publicly available COAs” → /coas/]
[INTERNAL-LINK: “MOTS-c product page” → /product/mots-c/]
Frequently Asked Questions About MOTS-c
Is MOTS-c natural or synthetic?
Both, depending on context. MOTS-c is naturally encoded in the human mitochondrial genome — your body produces it. However, the MOTS-c available for research purchase is synthesized in a laboratory using solid-phase peptide synthesis. The synthetic version replicates the exact 16-amino-acid sequence described in the original 2015 discovery paper by Lee et al. (PMID: 25738459).
What does MOTS-c stand for?
MOTS-c stands for Mitochondrial Open Reading Frame of the 12S rRNA-c. “Open reading frame” means a stretch of DNA encoding a peptide. “12S rRNA” is the specific mitochondrial gene where the instructions were found. The “c” differentiates it from other peptide sequences in the same gene region.
How is MOTS-c different from SS-31?
SS-31 is a synthetic peptide designed to target mitochondria from outside the cell. MOTS-c is naturally encoded by mitochondrial DNA and travels outward — from mitochondria to cytoplasm, nucleus, and bloodstream. They’re studied for different research questions: SS-31 for mitochondrial membrane integrity, MOTS-c for systemic metabolic signaling.
For research use only. Not for human consumption. MOTS-c is an experimental compound with no FDA-approved therapeutic applications. All information on this page is provided for educational purposes relating to laboratory and preclinical research.
[INTERNAL-LINK: “MOTS-c” → /product/mots-c/]
[INTERNAL-LINK: “Certificates of Analysis” → /coas/]